Advances in microbiome science are occurring faster than our ability to apply these insights within complex human systems. While research has dramatically expanded what can be observed within the gut, translating these discoveries into meaningful improvements in patient outcomes remains inherently difficult.
From Discovery to Clinical Application
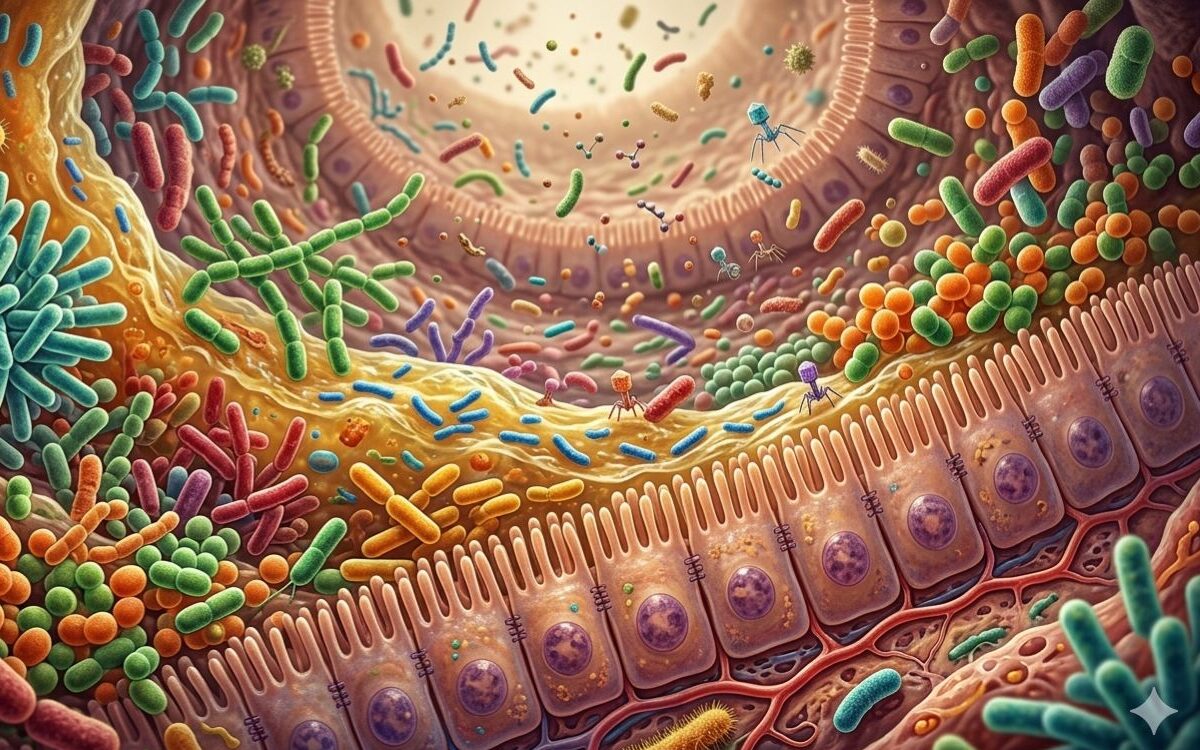
Over the past two decades, research into the gut microbiome has significantly expanded scientific understanding of human biology and opened new possibilities for clinical insight. Advances in sequencing technologies and metabolomic analysis have revealed that the trillions of microorganisms inhabiting the human gut actively participate in immune regulation, metabolic signalling and neurological communication.¹
Microbial metabolites influence inflammatory pathways, insulin sensitivity and neurotransmitter activity, while research on the microbiota–gut–brain axis highlights the close interaction between microbial activity and nervous system signalling.² ⁷
As these scientific insights have emerged, public awareness of gut health has grown alongside them. Direct-to-consumer microbiome testing and algorithm-driven dietary recommendations have entered everyday health decision-making as individuals seek personalised explanations for chronic digestive symptoms.⁸
Despite this rapid progress, a gap remains between microbiome discovery and its application in everyday clinical care. While microbiome data can reveal important microbial patterns, it rarely provides a clear roadmap for treatment when interpreted in isolation.³
For clinicians working with chronic digestive disorders, the challenge is often not how deeply we can analyse the microbiome, but how those insights are interpreted and applied within the complexity of individual physiology and lived experience.
Expanding Visibility in Digestive Health
Research has identified associations between microbial patterns and a wide range of disease states, including inflammatory bowel disease, metabolic syndrome and functional gastrointestinal disorders.⁶ These patterns often involve reduced microbial diversity and altered metabolite production.
In clinical and specialist settings, sequencing technologies allow clinicians to characterise microbial communities and establish baseline microbial profiles in the context of disease-associated patterns. Yet much of the microbiome literature that informs this work remains observational or laboratory-based, meaning these findings do not always translate into a clear understanding of what is occurring within individual patients.³ Consequently, microbiome data is often most useful as a marker of change, allowing clinicians to track shifts in microbial ecology alongside symptoms, dietary patterns and environmental exposures as treatment progresses.
Interpretation of microbiome data is further complicated by the dynamic nature of the microbiome itself. Stool sequencing provides only a snapshot of a highly dynamic ecosystem shaped by diet, medication exposure and immune activity, meaning that a single test result may reflect recent inputs rather than a stable representation of long-term microbial ecology.⁵
Individual variation adds another layer of complexity. Even among healthy populations, microbial communities vary substantially between individuals and are shaped by genetics, dietary patterns, lifestyle factors and ongoing environmental exposures.⁴ Similarly, individuals with comparable microbial profiles may experience very different symptom patterns due to differences in host physiology and regulatory processes.
For these reasons, microbiome findings require careful clinical interpretation before they can be meaningfully applied in patient care. Understanding the environment in which microbial communities exist therefore becomes central to interpreting microbiome data.³
Host Physiology as the Microbial Environment
The gut microbiome exists within a physiological environment shaped by digestive secretions, intestinal motility, mucosal immune activity and host regulatory processes.¹ These factors influence which microbes thrive, how microbial metabolites are produced and how the microbial ecosystem interacts with the host.
Digestive function, immune regulation and stress physiology shape the microbial habitat itself and strongly influence how individuals respond to dietary or therapeutic interventions.
Nervous system signalling forms a central component of this regulatory environment, as it coordinates many physiological processes including those that shape digestive function and host defence. Research describing the gut–brain axis has shown that stress responses can alter digestive function and intestinal permeability, thereby reshaping the microbial habitat itself.²
In individuals with long-standing digestive disorders, nervous system dysregulation is frequently observed alongside microbial imbalance. In practical terms, this often reflects changes in how well someone digests food, how reactive their immune system is, and the level of stress their nervous system is under.
When Biological Insight Meets Real Patients
The complexity of digestive health often becomes most visible when biological insight meets the lived reality of patients.
Individuals with chronic digestive symptoms frequently arrive after years of fluctuating health, repeated dietary interventions and multiple healthcare consultations. Many have experimented extensively with elimination diets, supplements or restrictive protocols in an attempt to regain control of their condition.
Over time these efforts can place the nervous system in a persistent state of vigilance around food and physical responses. Irregular meal patterns, restrictive eating and ongoing symptom monitoring can reinforce stress responses that disrupt digestive regulation and alter the microbial environment itself.
When nervous system dysregulation and digestive instability are present together, microbiome-targeted interventions may be only partially effective. In clinical practice, how stable someone feels around food and how settled they feel in their body often determines whether microbial interventions can be tolerated at all.
Progress often involves calming these regulatory patterns before meaningful microbiome change can occur. In many cases, restoring consistent nourishment, reintroducing dietary diversity and supporting nervous system stability must occur before sustainable microbiome change becomes possible.
Industry Expansion and the Interpretation Challenge
Rapid growth in microbiome testing has brought increasingly detailed microbial data into everyday health decision-making.
However, direct-to-consumer diagnostics and algorithm-driven dietary recommendations can generate highly detailed microbiome reports in the absence of appropriate professional interpretation. In clinical nutrition practice it is increasingly common for individuals to arrive with extensive microbiome reports but limited clarity about how the findings relate to digestive physiology or everyday eating patterns.
Without professional guidance, microbiome results are often interpreted in isolation, leading to unnecessary dietary restriction, excessive supplementation, misdiagnosis and undue concern. When presented without appropriate clinical context, increasing volumes of microbiome data can actually intensify vigilance around food and symptoms, reinforcing nervous system patterns that perpetuate gut imbalance and push recovery further out of reach.
Communication within the microbiome field also shapes public expectations about what microbiome testing can realistically deliver. Research findings often reach the public through media coverage and commercial platforms long before practical clinical guidance is established.
For many individuals, once microbial imbalance is identified, the expectation is understandably one of rapid correction. In reality, meaningful change in digestive systems that have been dysregulated for years rarely occurs quickly. Restoring stability typically requires gradual shifts in physiology, behaviour and microbial ecology.
Accurate communication about both opportunity and limitation, together with appropriate clinical interpretation, is therefore essential if microbiome science is to translate meaningfully into clinical care.
The Next Phase of Microbiome Medicine
The microbiome represents one of the most promising areas of modern biomedical research. Continued advances in sequencing technologies, metabolomics and microbial ecology are deepening understanding of host–microbe interactions.
However, digestive health operates as an ecological system shaped by differences in host physiology, regulatory processes and lived experience. As a result, stable microbial ecosystems and digestive balance are more often restored through sustained environmental support rather than isolated intervention.
The next phase of microbiome medicine therefore depends as much on integrative clinical interpretation as on further technological innovation. Bridging microbiology, nutrition science and patient-centred clinical practice will be essential if these discoveries are to translate into meaningful health outcomes.
Ultimately, progress will depend not only on deeper examination of microbial ecosystems, but on a more detailed and nuanced understanding of the individuals within whom those ecosystems exist.
Author Bio

Amanda Callenberg, Nutritional Therapist (BANT, CNHC)
Amanda Callenberg is a London-based Registered Nutritional Therapist (BANT, CNHC) specialising in chronic digestive disorders and IBS. Her clinical focus is on the interaction between nutrition, stress physiology, nervous system regulation and the gut microbiome in long-term digestive recovery. Her work integrates microbiome testing with nervous system-informed nutritional therapy to support people experiencing persistent digestive symptoms.
Website: https://amandacallenberg.com
LinkedIn: https://www.linkedin.com/in/amandacallenberg/