In the pharmaceutical industry, the statistics of failure are so familiar they have become clichés: it takes ten years and two billion dollars to bring a new drug to market, with a failure rate in clinical trials hovering near 90%. While much of this attrition is attributed to biological complexity, a significant, often overlooked bottleneck lies in the fundamental execution of pre-clinical research.
The crisis in drug discovery is not solely a crisis of biology; it is a crisis of methodological anarchy.
For decades, scientific research has been guarded behind the closed doors of labs and corporate R&D centers. Critical details are frequently treated as proprietary trade secrets or, conversely, are standardized so poorly that they become “tribal knowledge,” accessible only to a select few within a specific institution. This siloing of Standard Operating Procedures (SOPs) creates a fragmented landscape where reproducibility suffers, and vital research momentum is lost to unnecessary troubleshooting.
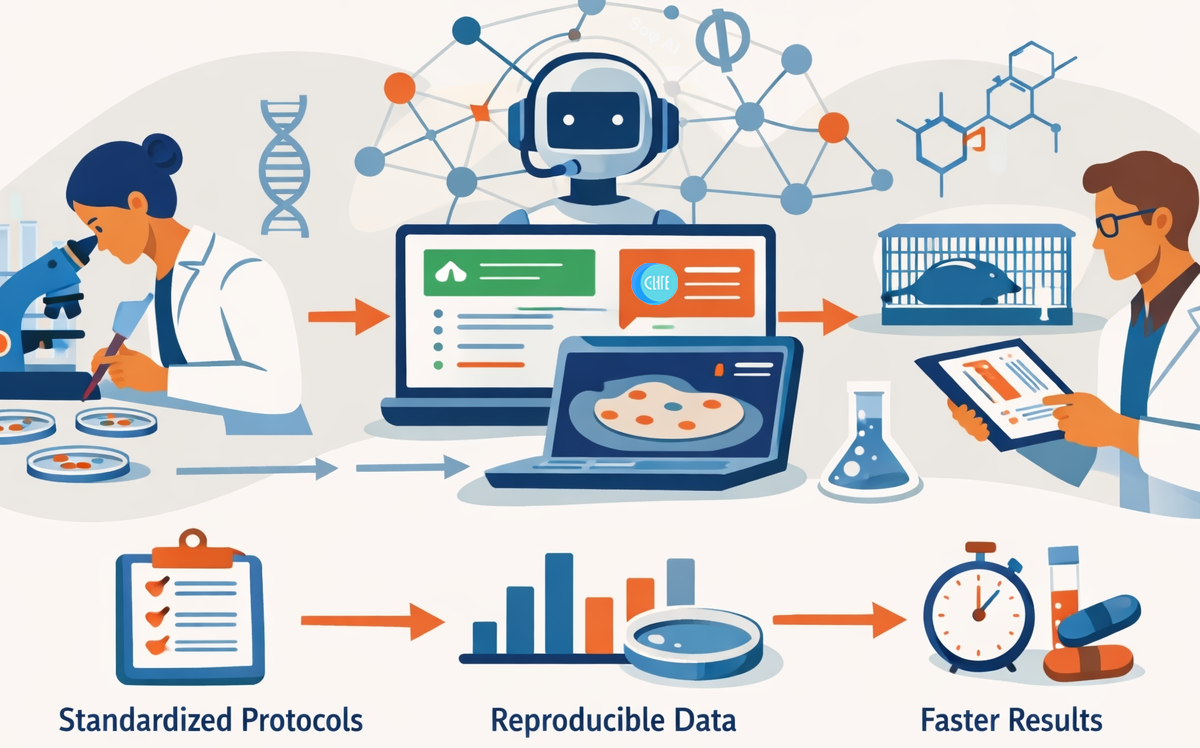
This is where one of the biggest developments in recent life sciences has become apparent: the emergence of artificial intelligence that does not merely analyze experimental data, but helps standardize entire lab workflows. These AI-powered laboratory assistants represent a paradigm shift in how we approach experimental standardization and data analysis in drug discovery pipelines.
The Standardization Challenge
Currently, a researcher tasked with a new assay might spend days scouring literature, often finding protocols that are vague or incomplete. They then struggle to adapt these protocols to their specific equipment. This friction slows down the drug discovery pipeline at its earliest, most critical stage: target validation.
Drug discovery relies heavily on standardized assays to evaluate compound efficacy, toxicity, and mechanism of action. Consider the scratch (wound healing) assay, a widely used method for studying cell migration in cancer research, regenerative medicine, and drug screening. While conceptually simple, outcomes are highly sensitive to protocol details-cell density, scratch technique, imaging intervals, and analysis thresholds.
Small deviations in these variables can dramatically alter results, leading to conflicting conclusions between labs. Manual image analysis further compounds the issue, introducing operator bias in measurements and interpretation.
This pattern is not unique to scratch assays. Similar challenges appear across toxicity screens, viability assays, phenotypic imaging studies, and mechanistic investigations.
Lab protocols aren’t one-size-fits-all, and procedures that work in one lab might fail in another due to differences in equipment, reagents, or cell lines. This variability creates several cascading problems:
- Inter-laboratory inconsistency: When a promising compound shows anti-metastatic properties in one laboratory’s scratch assay but fails to replicate elsewhere, is the compound ineffective, or was the assay poorly standardized?
- Delayed timelines: Researchers spend weeks optimizing protocols for their specific conditions-time that could be devoted to hypothesis testing and data generation.
- Knowledge silos: Successful protocol adaptations often remain within individual labs, never formally documented or shared.
- Subjective analysis: Manual measurements of cell migration introduce operator bias, particularly in edge detection and threshold setting, leading to non-reproducible quantification.
These issues compound throughout the drug discovery pipeline, from target validation through preclinical efficacy studies, ultimately contributing to the high attrition rates in clinical development.
A Shift Toward Intelligent Standardization
In recent years, a new class of artificial intelligence systems has begun to emerge, focused not just on analyzing experimental data but on supporting the structure of laboratory work itself. Rather than replacing scientists, these tools aim to encode best practices, adapt protocols to specific lab environments, and reduce the operational friction that slows research progress.
Instead of relying on scattered literature searches and informal guidance, researchers can increasingly turn to AI-driven workflow systems that:
-
Translate published methods into clear, step-by-step procedures
-
Adapt protocols to specific instruments and reagents
-
Preserve procedural knowledge in standardized digital formats
-
Apply consistent analytical approaches across experiments
The goal is not automation for its own sake, but reproducibility at scale.
AI-powered research assistants are beginning to address these challenges by functioning as centralized intelligence for laboratory operations. Rather than simply storing scientific knowledge, these systems understand the logistics of experimentation. By processing vast datasets of methodological constraints, they can generate tailored, step-by-step SOPs adapted to a researcher’s specific hardware and reagents. Several such platforms are working to unlock the “guarded” knowledge of the industry, aiming to provide more universal standards that enable a graduate student in a small biotech startup to execute an experiment with similar precision to a veteran in a top-tier pharmaceutical company.
Efficiency as a Catalyst for Discovery
The automation of tedious analytical tasks represents another significant advancement. In scratch assays, for example, analyzing cell images has traditionally involved manual tracing or utilizing legacy software that introduces human bias. Computer vision technology can now detect cell borders and calculate wound closure rates automatically. This automation offers several advantages:
Removing subjectivity: Data becomes strictly quantitative, eliminating the “human error” variable that often leads to false positives in drug screening.
Accelerating throughput: Researchers can process hundreds of images in the time it once took to process one, allowing for more rapid iteration of drug candidates.
By offloading prone-to-error tasks like calculation, protocol writing, and image analysis to AI systems, scientists are freed to focus on high-level interpretation and hypothesis generation; as these new tools can:
-
Shorten experimental setup time
-
Reduce methodological drift
-
Improve cross-team reproducibility
-
Generate cleaner datasets for downstream decision-making
Rather than relying solely on individual expertise, organizations begin to operate on continuously improving procedural frameworks.
Implications Across Drug Discovery
As intelligent standardization matures, its impact is expected across multiple stages:
Target validation becomes more reliable as assays are executed consistently across cell models and labs.
Compound screening benefits from faster turnaround and reduced analytical noise, improving lead prioritization.
Preclinical studies gain stronger reproducibility, supporting regulatory confidence and program continuity.
Ultimately, better operational rigor upstream translates into stronger scientific foundations downstream.
The Future of Research Execution
The next major acceleration in drug discovery may not come solely from new algorithms predicting molecules or structures, but from transforming how experiments themselves are run.
By systematizing protocols, preserving procedural knowledge, and removing avoidable variability, AI-driven workflow systems are turning laboratory execution from an artisanal process into a scalable, reproducible engine of discovery.
As research grows more complex and interdisciplinary, the ability to standardize without sacrificing flexibility will likely become a defining competitive advantage in life sciences.
The future of drug development will be shaped not only by what scientists discover-but by how reliably, efficiently, and consistently those discoveries are made.
Author Bio:
Mojtaba is a Product Development Engineer at Cornell University and the founder of CLYTE, a startup dedicated to standardizing laboratory workflows through intelligent automation. With a degree in Computer Science, he currently developed “Soφ AI” a framework for experimental reproducibility, by providing intelligent SOPs, troubleshooting, and data analysis for life science research. His work centers on utilizing AI and computer vision to eliminate methodological variability in preclinical research.